Grids
There are many different types of grids used by ocean models, such as latitude-longitude grids, tiled rectilinear grids, triangular or hexagonal grids, and adaptive unstructured grids. An observational campaign could have stations at arbitrary locations.
neutralocean can handle any dataset of vertical casts: you just have to specify which pairs of casts are adjacent. We think of the horizontal grid as a graph: a collection of nodes and edges. Each node is a vertical cast (which could be ocean or land). Two nodes are joined by an edge when the two vertical casts are adjacent.
Example. Suppose we took 4 casts from the ocean, with three in a triangle and a fourth off to the side (as in the run_4casts.py Test Example):
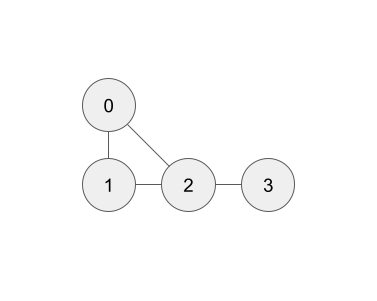
The nodes (vertical casts) are labelled 0, 1, ... N-1, where N is the number of casts (here N = 4).
The order doesn’t really matter, but once it’s chosen we then specify the edges (pairs of adjacent casts) as a 2D array of shape (2, E). Here, E = 4 and edges = np.array([[0, 0, 1, 2], [1, 2, 2, 3]]). Alternatively, edges can be a tuple/list of length 2, with each element a tuple/list of length E: edges = ((0, 0, 1, 2), (1, 2, 2, 3)). What matters is, letting a = edges[0] and b = edges[1], that the nodes a[i] and b[i] are adjacent, for each i = 0, ..., E-1.
For a rectilinear grid (such as a lat-lon grid), most casts are adjacent to four other casts. We can label the casts 0, 1, ..., N-1 starting in the south-west and going first across longitudes and then across latitudes, ending in the north-east. The order doesn’t really matter, but must be specified.
Example. A 4 x 3 grid with N = 12 that is periodic in longitude (x-axis) but not in latitude (y-axis):
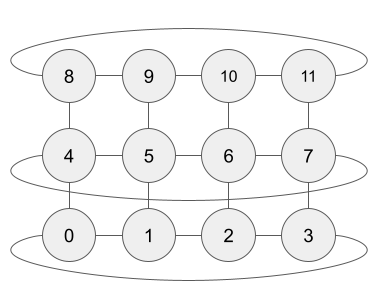
Note
The order of casts is determined by your data is stored in memory. Suppose Salinity and Temperature are presented as 3D arrays of size (nj,ni,nk), which are the number of grid points in the latitudinal, longitudinal, and vertical dimensions, respectively. The data is actually stored in memory as a long 1D array with the first nk elements being data from the “first” cast, the next nk elements being data from the “second” cast, etc. This orders the casts. Ideally, the vertical dimension is last (varying fastest in memory); if not, you can use the vert_dim argument and neutralocean will internally reorder your data to be so.
The pairs of adjacent casts can be encoded as
edges = (
(0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 4, 5, 6, 7, 8, 9, 10, 11),
(3, 0, 1, 2, 7, 4, 5, 6, 11, 8, 9, 10, 0, 1, 2, 3, 4, 5, 6, 7),
)
# <--------- x connections ----------->, <--- y connections --->
If the grid also gives geometric information, we should give it to neutralocean.
If casts m and n are adjacent,
let \(\Delta_{m,n}\) be the distance between cell centers of casts m and n, and
let \(\Delta^\perp_{m,n}\) be the distance of the face between the cells of casts m and n.
This geometric information is encoded as two E-length lists, dist and distperp respectively, given in the same order as edges. E.g. dist[i] is the distance between casts edges[0][i] and edges[1][i].
This geometry for the 4x3 example can be illustrated as follows (not showing all geometric info):
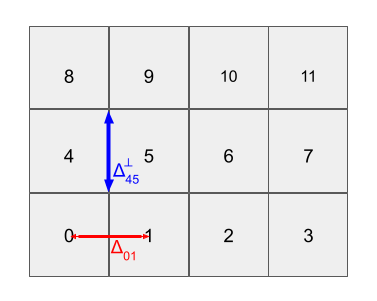
The lists edges, dist, and distperp are passed to neutralocean as a dict named grid (see Potential Density Surface for its docstring).
To ease building these lists, we provide build_grid functions for various grid types.
See the Grids API.
For the first generic graph example, we’d use neutralocean.grid.graph.build_grid.
For the lat-lon example, we’d use neutralocean.grid.rectilinear.build_grid. See the OCCA example.
A close cousin of rectilinear grids (like lat-lon) is a tiled rectilinear grid, in which there are several (square) rectilinear grids that are placed next to each other, such as the lat-lon-cap used by ECCOv4r4. For these grids, use neutralocean.grid.xgcm.build_grid to build the grid dict. This uses xgcm to handle the tiling, with face connections specified in the xgcm way. See the ECCOv4 example.
For a tripolar grid such as ORCA used by NEMO, we’d use neutralocean.grid.tripolar.build_grid. See the ORCA example.